Introduction to 
cellXpress is a biologist-friendly image analysis and visualization software tool specifically designed for multiplexed fluorescence (MxF) cellular and tissue images. Unlike most other existing image analysis tools, it does not require the users to have any prior programming experience.
cellXpress is a highly efficient tool written in C++ and optimized for modern processors. The new cellXpress 2 has several new features specifically designed for MxF tissue images.
 Marker-set architecture
Marker-set architecture
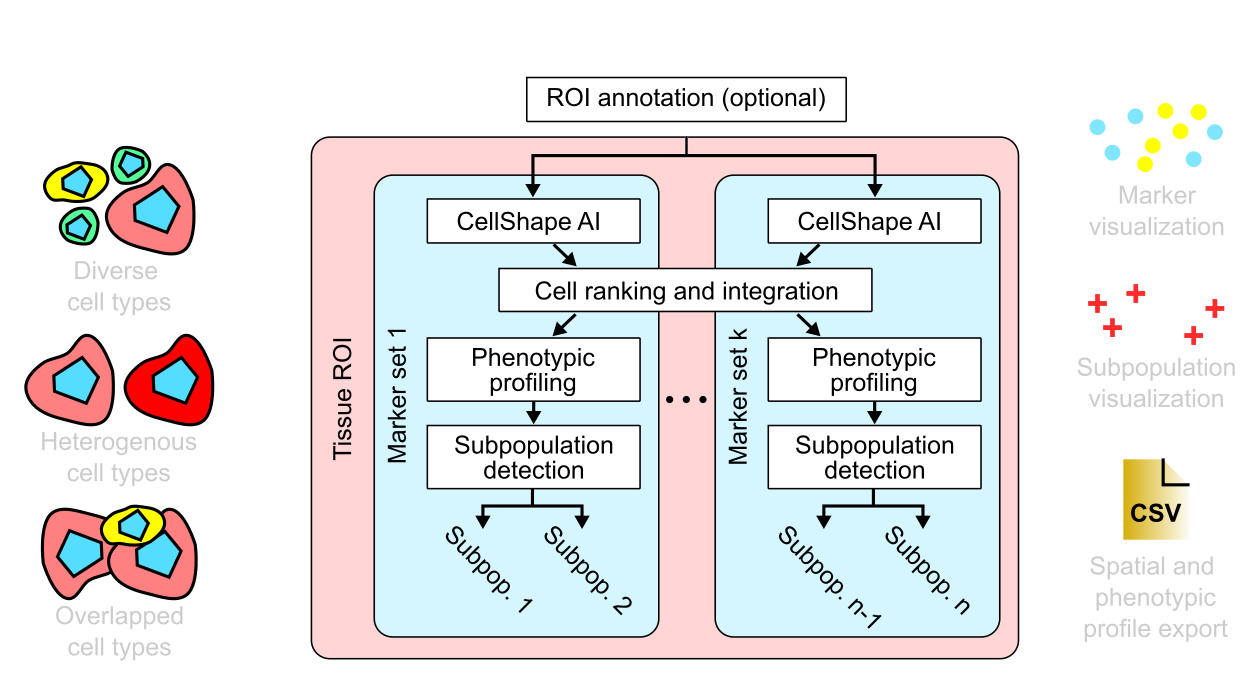
cellXpress has a unique “marker-set” processing and integration architecture that allows for modular selections and configurations of image processing and cell/subpopulation detection algorithms based on different combinations of markers.
Thus, different processing settings can be flexibly configured for different subsets or combinations of markers, which usually reflect different cell types or states, within the same tissue images.
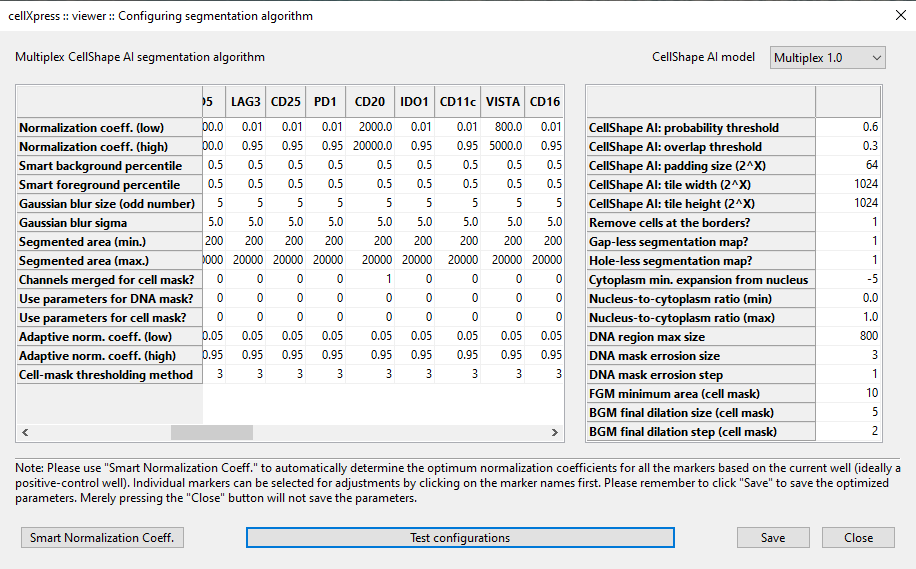
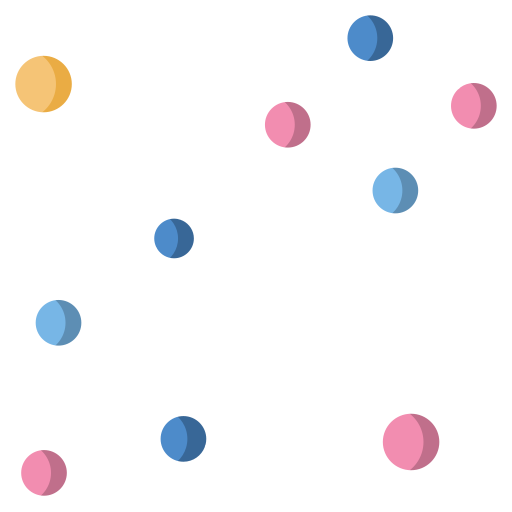 CellShape AI Segmentation
CellShape AI Segmentation
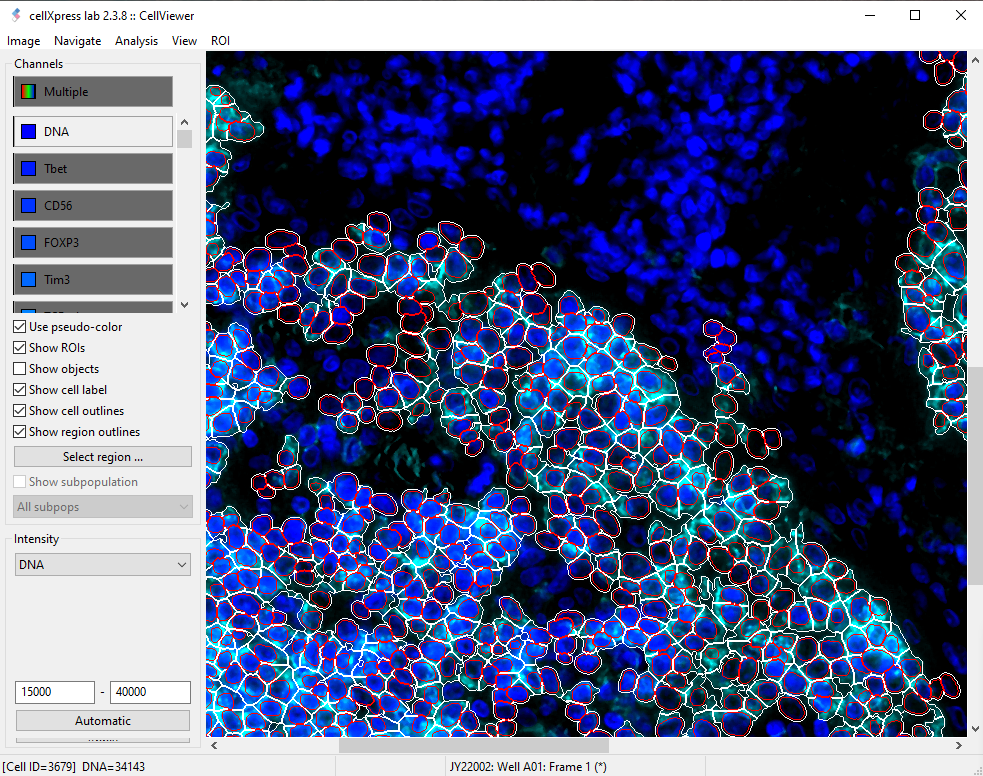
cellXpress contains a built-in and DNN-based cell detection algorithm, “CellShape AI” (CSAI), that can robustly identify nuclear and cellular boundaries even on cells with heterogonous staining levels, without requiring any additional software setup, nor custom scripting.
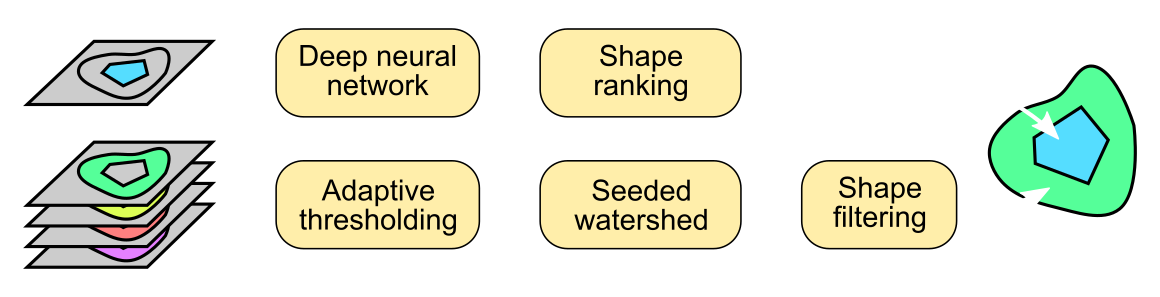
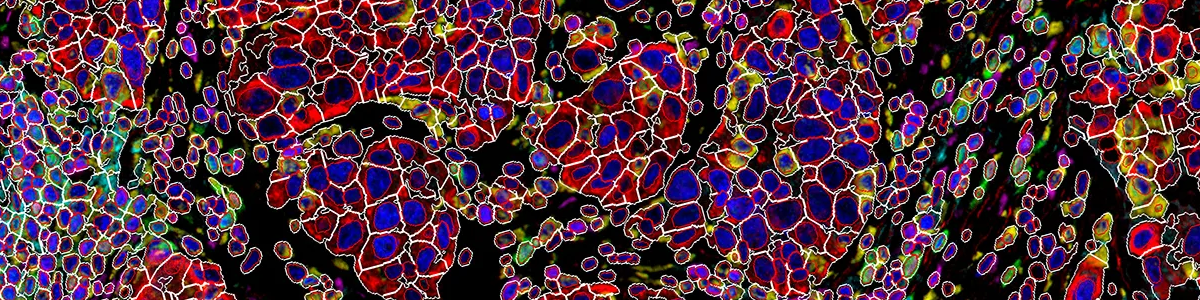
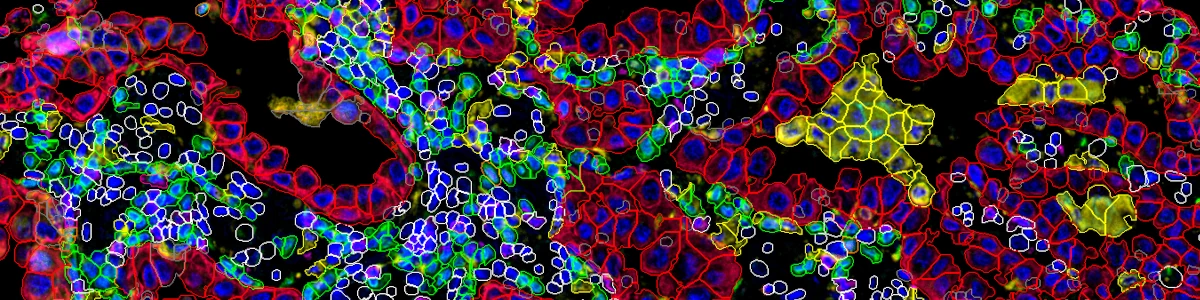
Furthermore, our unique marker-set processing architecture also enables CSAI to be flexibly applied to different combinations of markers within the same images, and thus making cellXpress to be currently the only tool that can detect overlapped cell types and quantify their boundaries.


 Marker-set architecture
Marker-set architecture
 CellShape AI Segmentation
CellShape AI Segmentation


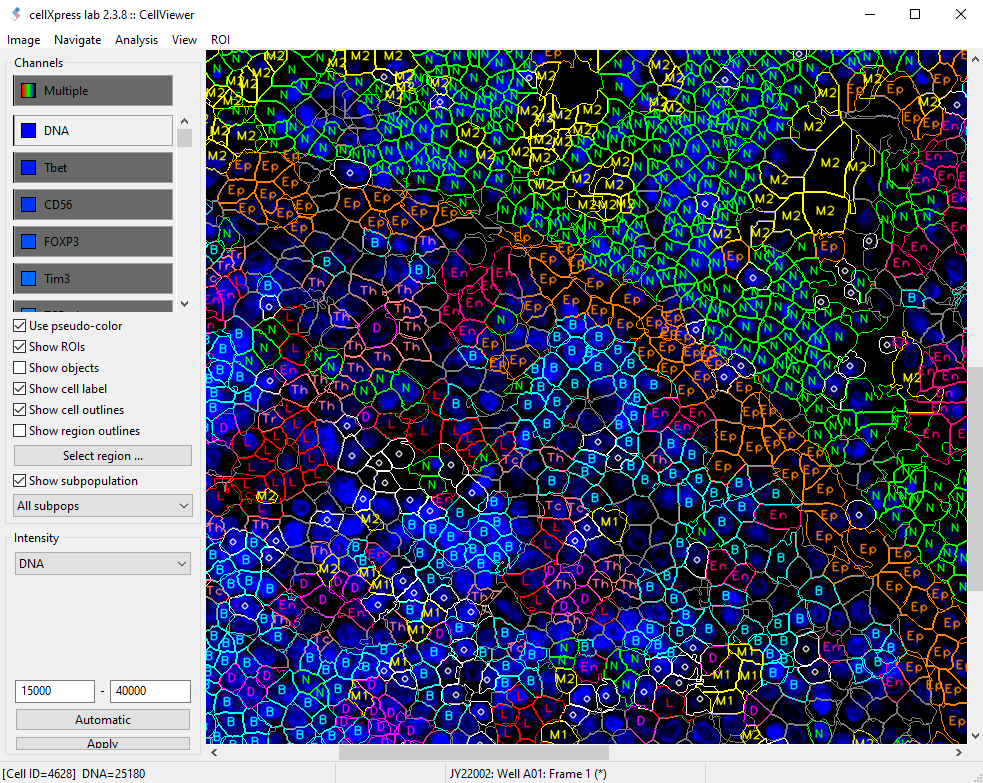
 GUI for Hyperplexed Images
GUI for Hyperplexed Images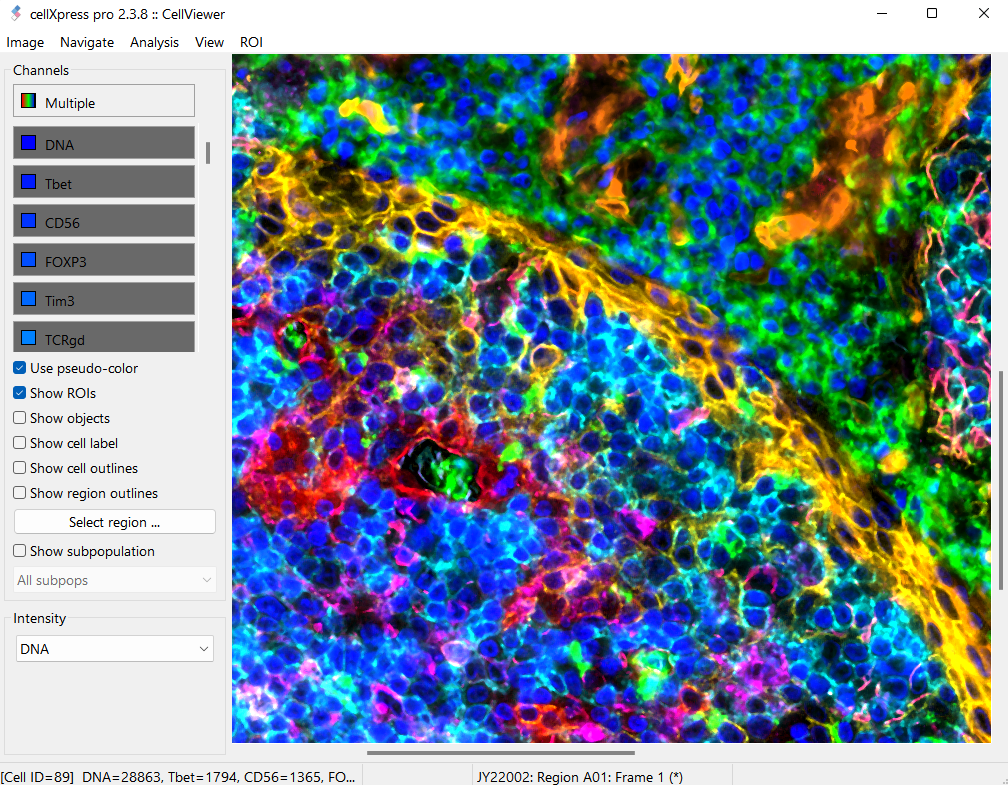
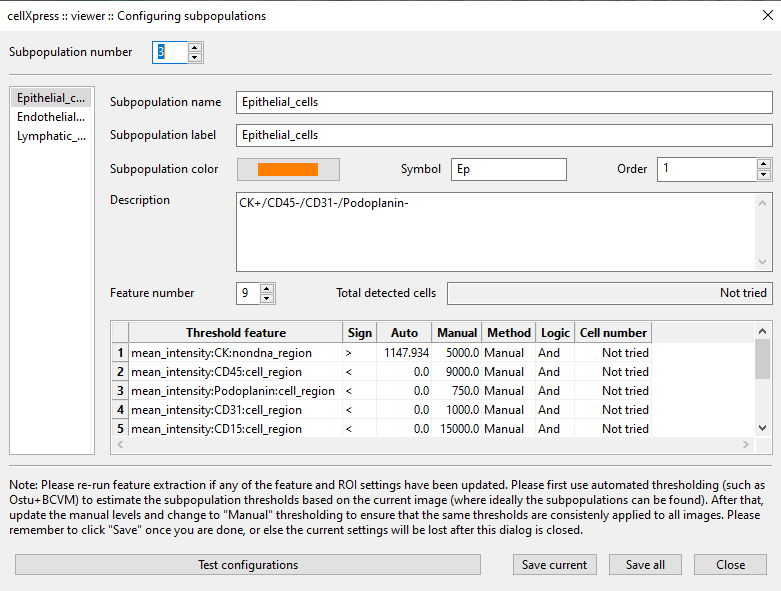
 Subpopulation identification
Subpopulation identification